ARS for Pathogen Genomic Center of Excellence
Update: this position has been closed.
The Grubaugh Lab at the Yale School of Public Health (YSPH) is seeking a Associate Research Scientist (or postdoc with relevant experience) with a background in pathogen genomics and bioinformatics to help lead Yale’s academic partnership to the Pathogen Genomic Center of Excellence (PGCOE). The Massachusetts Department of Public Health (MDPH) selected the YSPH and five other partners to make up its new PGCOE, one of five such centers recently launched by the U.S. Centers for Disease Control and Prevention (CDC). With $25 million in funding over five years, the center’s goal will be to improve technologies and coordinated processes for systematically monitoring infectious diseases and emerging variants, enabling the U.S. to better prevent and respond to outbreaks.
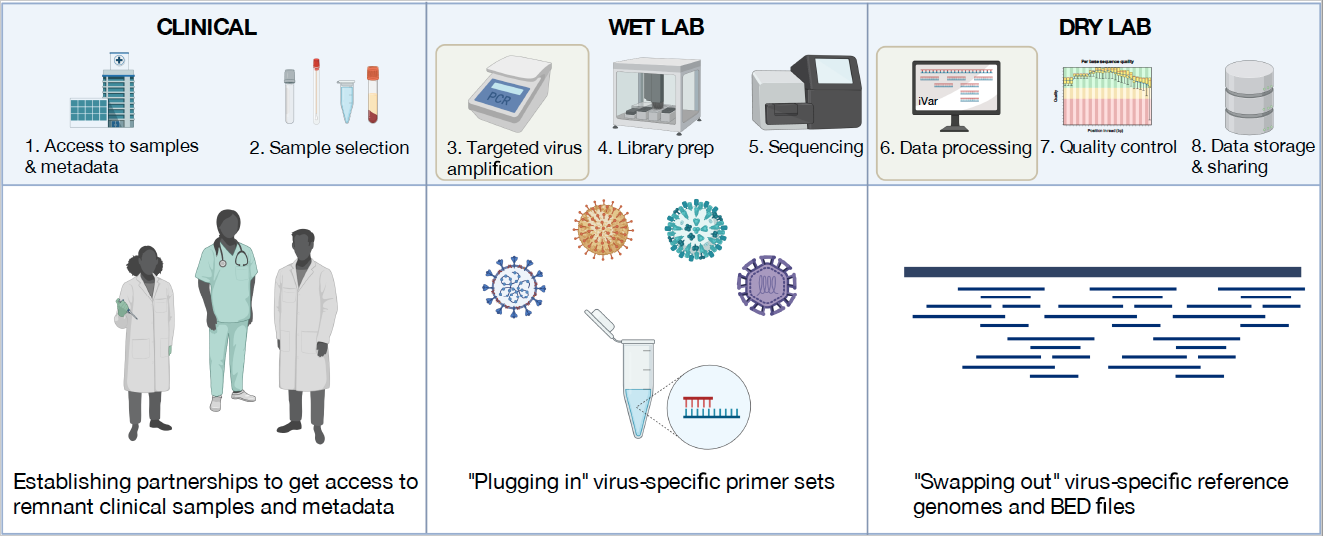
The position would entail leading a team of postdocs, research associates, and students to develop and validate new sequencing assays and bioinformatic pipelines, work with public health and clinical laboratories to translate the technology, and to respond to outbreaks. The CDC grant, subcontracted from the MDPH, is for five years. The first year will focus on developing a panel of amplicon-based sequencing strategies for respiratory viruses and exploring the use of highly multiplexed PCR for sequencing respiratory bacteria. The Grubaugh Lab will also collaborate with the Yale New Haven Hospital diagnostic labs for clinical validation and with the other academic partners (The Broad Institute, Boston University, Fathom Information Design, Massachusetts General Hospital, and Theiagen Genomics) to implement the approaches and discuss future directions. Future avenues of this grant may include: (1) wastewater sequencing, (2) broad-spectrum sequencing approaches, (3) genomic epidemiology of regional pathogens of interest (e.g. arboviruses), and (4) implementing routine genomic surveillance. The applicant would directly interact with state public health labs, academic partners, and the CDC.
Job requirements
The ideal candidate will have:
- PhD in microbiology, bioinformatics, or a related field
- 3+ years experience with library preparation and sequencing platforms
- strong bioinformatic skills: at a minimum, able to install, use, and edit bioinformatic programs
- good oral and written communication skills
- ideally: 3+ years of postdoc experience (though will consider less with extensive experience above)
Start date, duration and salary
Ideally, the candidate would be able to start between January 1 and April 1, 2023, though later start dates may be considered. The project is for 5 years, and includes the possibility for promotion and extension. Starting salary for an Associate Research Scientists (with at least 3 years of postdoc experience) is competitive and negotiable. Starting salary for a postdoc is $60,000 (i.e. directly upon completing a PhD), with increases based on relevant experience. Salary will also include competitive benefits, and support for travel to conferences will be provided. In person and hybrid work is available.
Working with us
Good science isn’t done in a vacuum. Our projects are primarily team-based, both within our Yale group and with our many external collaborators. We strive to create a diverse and inclusive atmosphere and bring members into our team with unique and complementary skill sets. We are flexible to the challenges in our lives (e.g. time off, remote work), try to have fun, and celebrate successes. Read more about our team here.
Yale University is an Affirmative Action/Equal Opportunity Employer. Yale values diversity among its students, staff, and faculty and strongly welcomes applications from women, persons with disabilities, protected veterans, and underrepresented minorities.
Want to join our team?!
Interested applicants should submit a curriculum vitae, a 1-2 page statement of research interests describing your qualifications, and contact information for three references to Nathan Grubaugh ([email protected]). Want to learn more first, just send us an email!
~Cheers,
Nate
